Structural biology is an inherently visual science. By examining the 3D structures of biomolecules, researchers gain insight into their functions and determine how they work. As a sighted person, J.H.-P. didn’t give this fact much thought — until a blind student joined her team.
O.S., who has been blind since birth, enrolled to do a chemistry degree at the University of Delaware in Newark in late 2019. She was fascinated by the idea of conducting computational chemistry research, but was unsure of the process and the potential challenges that the work would present with regard to accessibility. Surely working with hazardous chemicals, fragile glassware and complicated equipment would pose substantial barriers for her and her guide dog, Ripple. Then, a Research Experiences for Undergraduates programme for chemists with disabilities provided O.S. the opportunity to engage in research in an environment that was conscious of her abilities and needs. The programme’s coordinators connected O.S. with J.H.-P., whose computational structural-biology projects offered an alternative to benchwork.
J.H.-P., then in her first year as a faculty member at the university, welcomed the challenge of approaching her research from a new perspective. With O.S., she enlisted C.Y., an assistive-technology specialist and assistant director of disability support services on campus. Motivated by the conviction that students with disabilities should have equal access to science, C.Y. was excited to collaborate with them.
These tools help visually impaired scientists read data and journals
We set out to develop a research programme that would give O.S. an experience to match that of her peers who aren’t blind. What followed was an exciting period of exploration and development that resulted in a publication, a poster at a national conference and a toolkit to help other researchers with low vision to interact in a field that they might not have realized was open to them.
Here’s how it happened.
Science-speak
Our first step was to investigate the assistive technology O.S. was already using, which included screen readers and refreshable Braille displays. Screen readers are programs that use synthesized speech to convey the information on a computer screen, allowing users to access digital content by listening, Braille displays use an array of mechanical pins that rise and lower to form Braille characters, enabling users to read text shown on a screen through touch. Once properly configured to operate in a scientific computing environment — for instance, to work on the Linux operating system — these tools facilitated many practical aspects of O.S.’s research.
Computational research is typically done using a text-based interface known as the command line. Researchers can use the command line to issue instructions to the computer, create and edit text files, write code and submit computing tasks for processing. By converting the text that she entered and received from the computer terminal into audio, O.S.’s screen reader gave her full access to this textual interface. Given that most high-performance software in computational chemistry, biophysics and structural biology include a text-only mode, she could download and manipulate the atomic coordinates of biomolecular structures, prepare the files needed to simulate their motion, run those simulations and analyse the results.
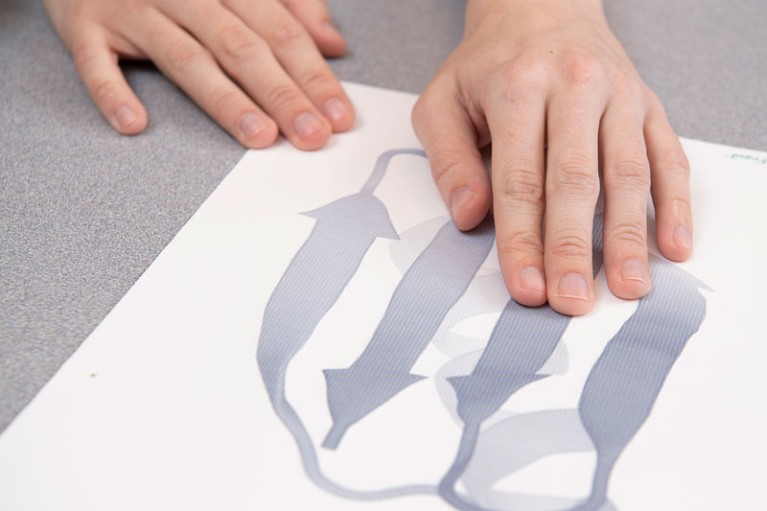

Raised images (known as tactile graphics) that are created on special type of paper are a useful tool for analysing 3D structures of proteins.Credit: Kathy F. Atkinson/University of Delaware
Occasionally, command-line information would appear in a format that was confusing when read aloud by the screen reader, such as tables. In those cases, J.H.-P. wrote code to automatically convert the information into a more linear format, for instance by transforming tabular data into short sentences.
The Emacs text editor, which can be controlled using keyboard short cuts or textual commands, along with the speech and audio interface, Emacspeak, allowed O.S. to quickly navigate and interact with the contents of text files. The Braille display was useful for coding and scanning through text in files or on the command line, particularly when the screen reader’s voice was unclear. Simple plots could be interpreted through sonification software, which transforms data into sounds, providing an auditory rather than visual form of analysis.
But what about the biomolecular structures? O.S. still needed a way to explore the complex 3D architecture of biomolecules that did not depend on sight. 3D-printed models were an obvious solution and proved useful for conceptualizing protein folds and secondary structures. Yet, these models took hours to produce, and were limited in the amount of structural detail they contained. They also could not convey the motion predicted by biomolecular simulations.
Chemical modelling with a sense of touch
To evaluate alternative strategies, we relied on C.Y.’s expertise in tactile graphics. Tactile graphics are raised images projecting from a flat background, which are produced by embossing or using a type of paper that swells with the application of heat, such that printed pictures rise up off of the page. Together, O.S. and J.H.-P. wrote a plug-in (called TactViz) for the widely used Visual Molecular Dynamics (VMD) software to depict proteins as cartoon diagrams, shaded by distance from the viewer. Such shading results in tactile graphics in which the spatial proximity of structural features to the viewer correlates with the elevation of the image, meaning that regions of the protein that appear closer to the viewer swell higher off the page. O.S. was able to discern secondary and tertiary protein structures from these graphics, which were quick and inexpensive to generate. We found that tactile graphics also provided a great way to create accessible plots, especially in cases in which the number of data points overwhelmed the sonification software.
Tactile dynamics
Of course, printed images cannot be rotated and manipulated as on-screen structures can, limiting O.S.’s ability to thoroughly examine complex biomolecules. However, we were then introduced to the Graphiti, a multi-line refreshable tactile display manufactured (and generously loaned to us) by Orbit Research in Wilmington, Delaware. When connected to O.S.’s computer, the Graphiti display acted as a monitor that was accessible by touch rather than sight, mirroring images on the screen with an array of mechanical pins, the heights of which adjusted automatically in response to changes in brightness. Our TactViz protein representations, already optimized for tactile graphics, displayed clearly on Graphiti, finally providing O.S. with interactive access to her biomolecular structures and, to some extent, their simulated motion.

Olivia Shaw’s three-year-old guide dog, Ripple.Credit: Kathy F. Atkinson/University of Delaware
Combining these assistive technologies with software tools that she developed to print text-based descriptions of the structural features of proteins and their spatial relationships, O.S. was finally positioned to conduct research on a similar level as her peers who aren’t blind. In one year, O.S. had collected data using a national supercomputer, submitted an accessible poster on her project to the American Chemical Society national meeting in 2020 (which was held virtually) and submitted a manuscript1 to the Journal of Science Education for Students with Disabilities.
Having lived her life in a world that was not designed with her circumstances in mind, O.S. was no stranger to devising workarounds to achieve her goals. Such flexibility and adaptability are essential skills for any researcher, and these qualities propelled her success in the lab. Also crucial to our collaboration was J.H.-P.’s and C.Y.’s responsiveness to O.S.’s stated needs, rather than their assumptions.
Through respectful communication, patience and outside-the-box thinking, we leveraged our combined expertise to develop tractable solutions for inclusivity that have impact beyond the experience of a single student. The community of blind and partially sighted people represents an untapped talent pool in science, technology, engineering and mathematics (STEM) fields, but tools designed with accessibility in mind can help researchers across multiple disciplines, regardless of visual acuity or limitations. We hope that by sharing our experience, we can motivate others to participate in training researchers who are blind or have low vision, as well as to consider their unique abilities and needs when developing technologies.